Tjelvar S.G. Olsson, Antony M.E. Churchill, William R. Pitt, John E. Ladbury & Mark A. Williams (2012). ProACT2 - Analysis of solvent accessibility, cavities and contacts in proteins and their complexes. Submitted
Mark A. Williams, Julia M. Goodfellow & Janet M. Thornton (1994). Buried waters and internal cavities in monomeric proteins. Protein Science 3, 1224-1235.
ProACT2
Analysis of protein accessibilities, cavities and contacts
Given a PDB format file containing one or more macromolecules, ligands and water molecules, PRO_ACT will;
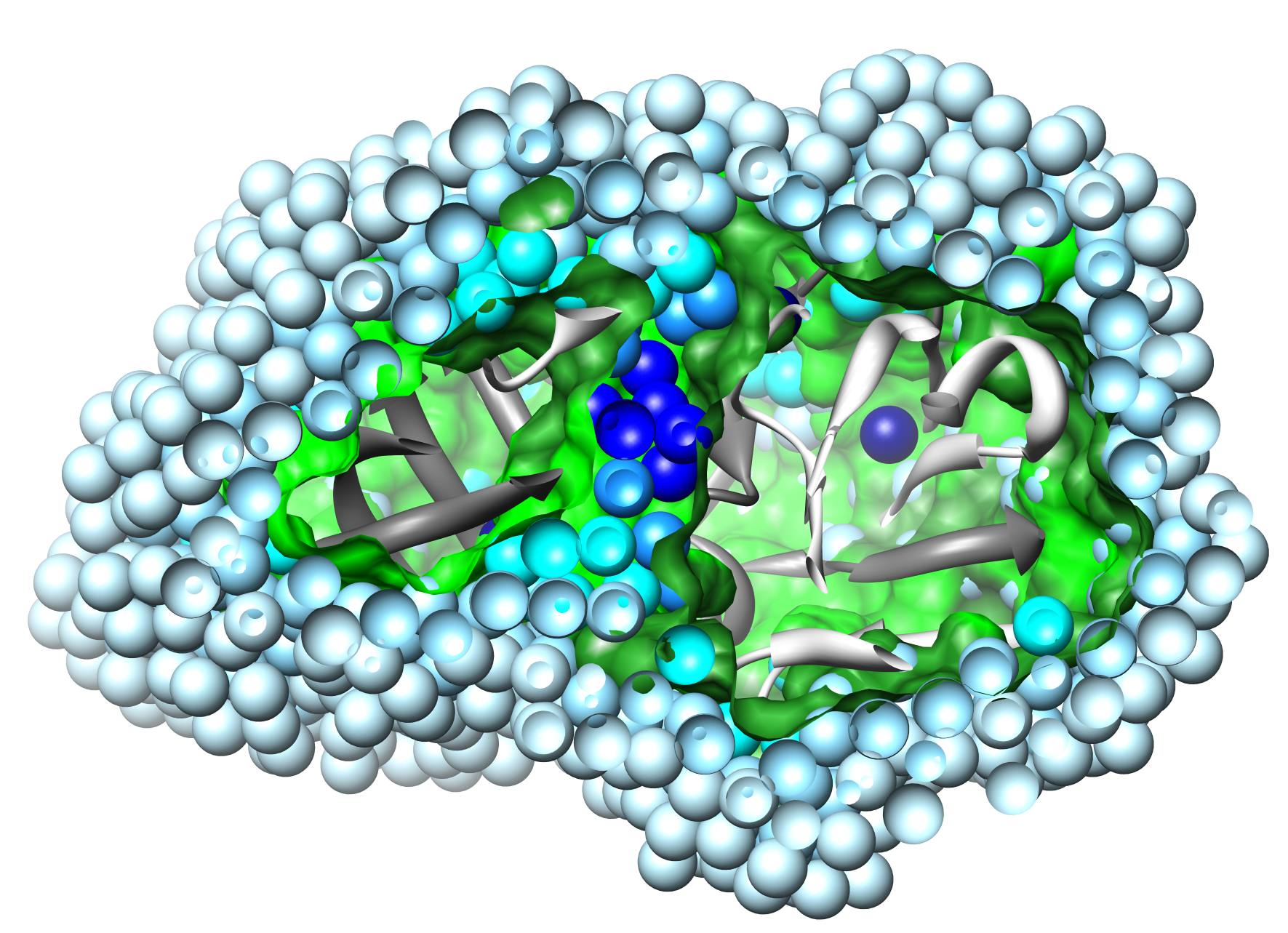
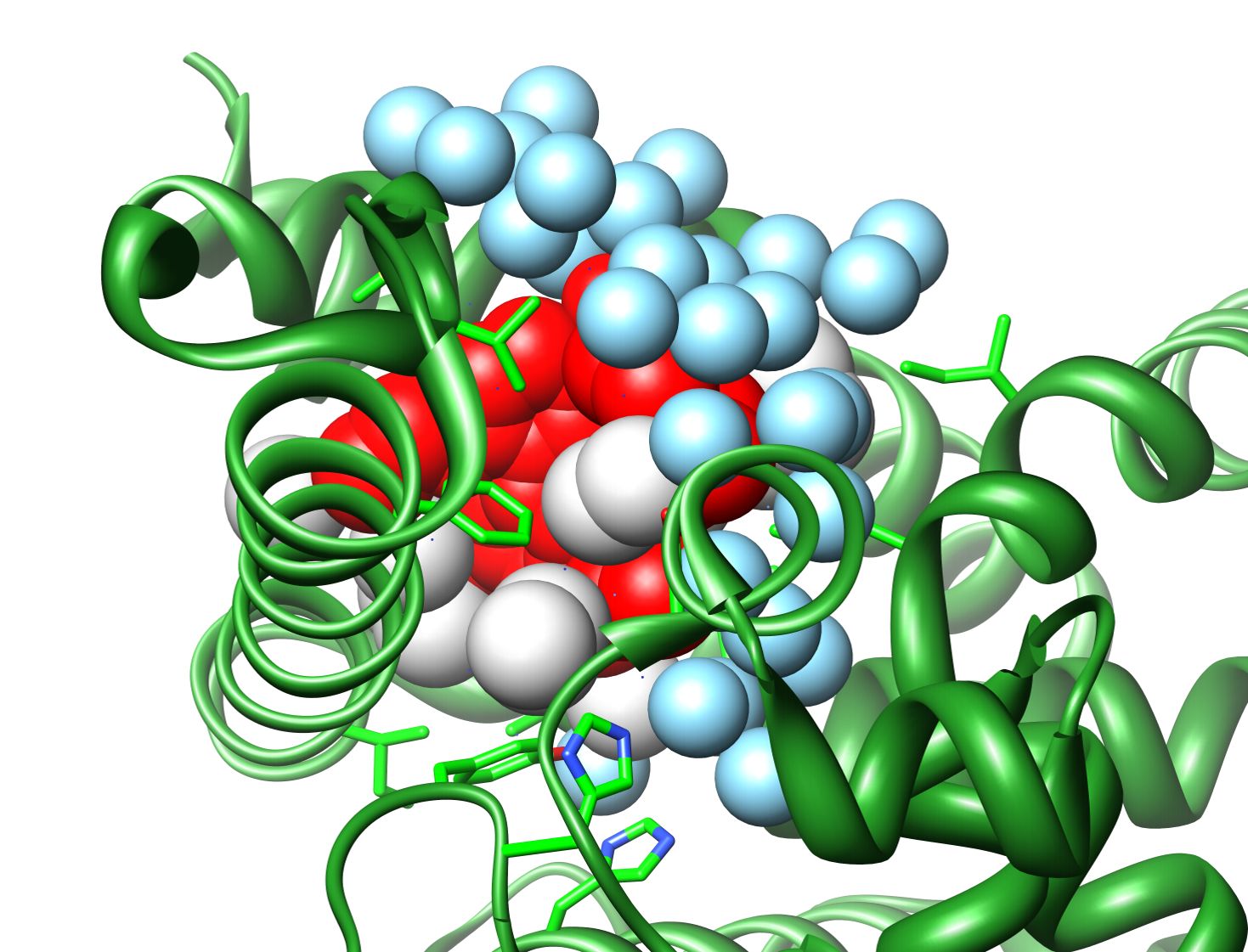
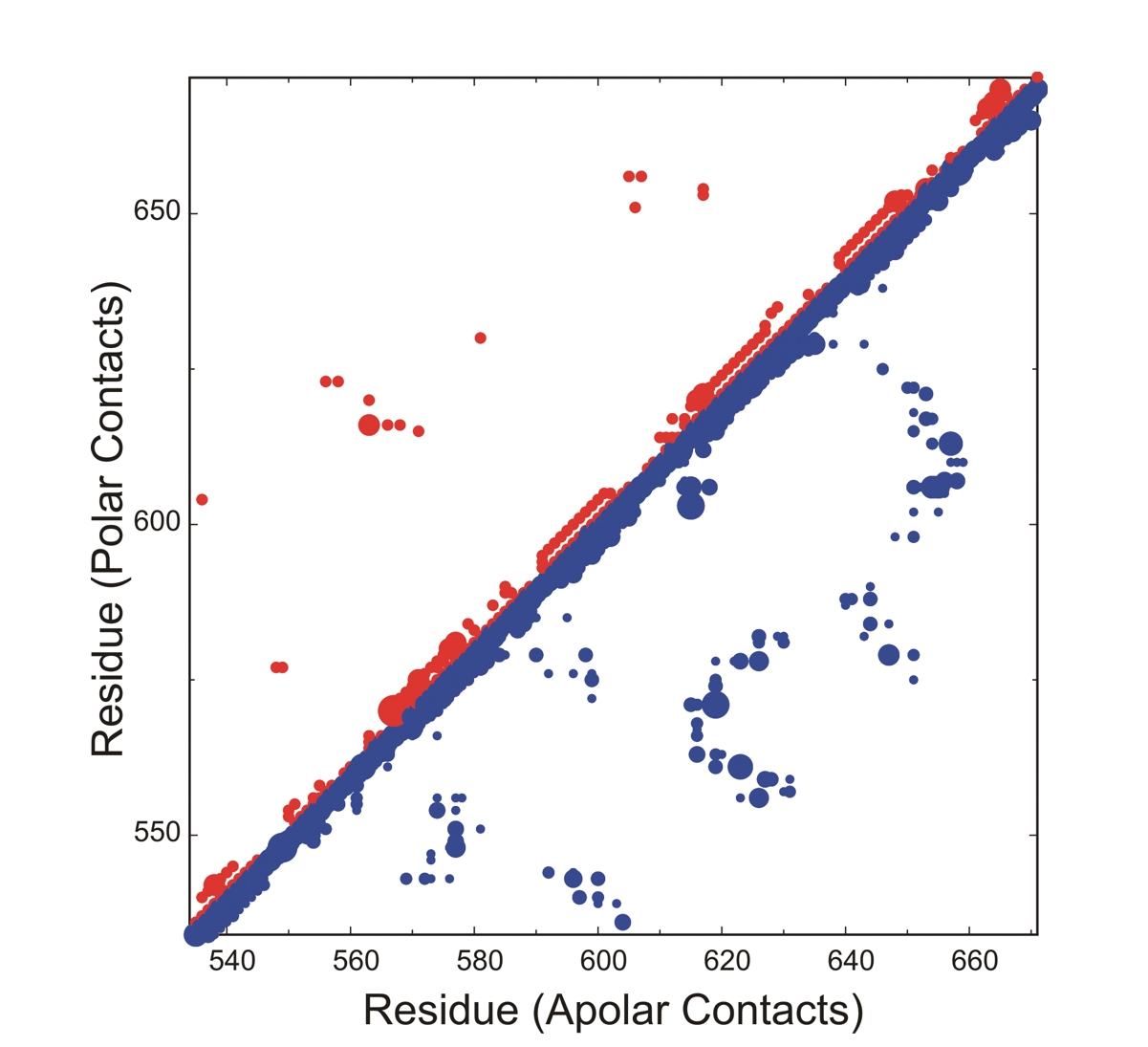
This new version of the software written in Python is based on the original PROACT algorithm for analysis of cavities and hydration, but functionality is greatly expanded, particularly in terms of its ability to deal with complexes and in the variety of output options. Some significant improvements have also been made to the robustness of the analysis (see manual for details). Atom-atom and residue-residue contacts are now avialable within and between components of a complex. Solvent accessibilities of proteins and ligands are also output in a manner familiar from other programs. The definitions of accessibility and cavity which are used by the software are consistent with a particular definition of contact, which is applied to atom-atom, atom-water and atom-cavity interactions. These definitions are rather different to those which are used by software based on the Lee and Richards/Connolly solvent accessible surface. The definitions and their rationale are discussed in the standard reference to the algorithm (Williams et al., 1994 - see box).
Download
The software is written in Python and should run on any common OS provided an appropriate python installation is available. The software can be downloaded here. In dowloading the software you agree to be bound by the terms of the Creative Commons licence under which it is released . The software is only licenced for non-commercial use. Any commercial users should contact the authors to discuss their requirements.
The manual is also included as part of the download.