Squid Photoreceptors
Helen Saibil
The retinal photoreceptors of the squid and other cephalopods
are composed of hexagonally packed cylinders of membrane (microvilli)
that contain the visual pigment, rhodopsin, along with the other
signal transduction proteins such as Gq and phospholipase C. The rhodopsin
is expected to have an ordered arrangement in the microvillar
membranes, because these animals can detect the plane of polarization
of polarized light. We
studied the signal transduction pathway in this system
using biochemical methods, and the rhodopsin structure by EM of
2D crystals.
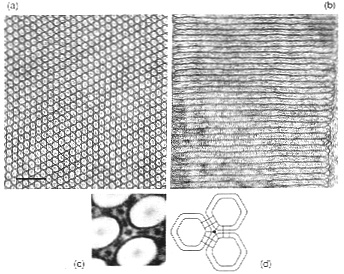
(a) Transverse and (b) longitudinal electron microscope sections
of photoreceptor microvilli from the squid retina. (c) Substructure
of the microvillar membranes. Image processing was used to show
2-fold and 3-fold structures linking the microvilli, suggesting
that these invertebrate photoreceptors have a more ordered structure
than bovine ROS membranes. This organization may provide the alignment
of rhodopsin molecules needed for discrimination of polarized
light by these animals. (d) Diagram of the membrane structures
seen in (c).
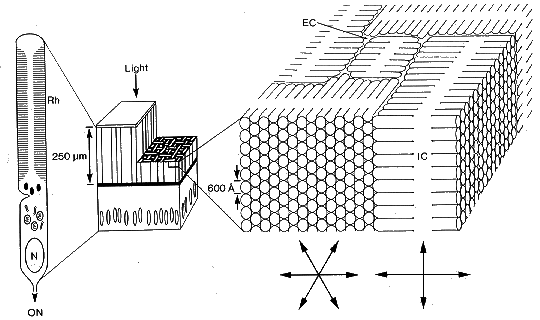
Schematic diagram of a slice of squid retina (centre) with expanded
areas showing the structure of a retinula cell (left) and the
arrangement of the photoreceptor membranes in the rhabdomes (right).
The photoreceptor layer of the retina (centre, vertical stripes)
faces the incoming light and the cut-away region shows the appearance
of the receptor mosaic in transverse section. The ribbon-like
outer segments of the photoreceptor cells weave together into
a retinal mosaic with the microvilli running in two orthogonal
directions (right). (Rh) rhabdome, (ON), optic nerve, (EC), extracellular
space, (IC), intracellular space.
This work was published in Saibil (1982) An ordered membrane cytoskeleton
network in squid photoreceptor microvilli. J. Mol. Biol. 158, 435 456,
and in
Saibil & Hewat (1987) Ordered transmembrane and extracellular structures
in squid photoreceptor microvilli. J. Cell Biol. 105, 19 28.
Rhodopsin structure
Our 2D crystals of squid rhodopsin led to an 8 Å projection
structure, published in:
Projection structure of an invertebrate rhodopsin.
Davies, A, Schertler, GFX, Gowen, BE & Saibil, HR (1996) J.
Struct. Biol. 117, 36-44,
and a 3D density map:
Three-dimensional structure of an invertebrate rhodopsin and basis for ordered
alignment in the photoreceptor membrane. Davies, A, Gowen, BE, Krebs, AM,
Schertler, GFX & Saibil, HR, (2001) J. Mol. Biol. 314, 465-473.

Projection structure of C-terminally truncated squid rhodopsin. One unit
cell is outlined and symmetry
elements are shown (space group p2221, a = 44 Å, b = 131
Å). Each unit cell contains 4 rhodopsin molecules. Contours
below the mean value are shown as dotted lines.
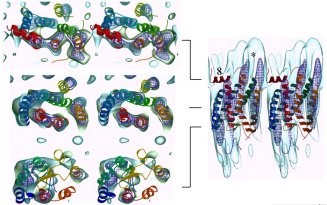
Sections and side views of two adjacent rhodopsin molecules in the lattice.
The squid rhodopsin density map is shown as a transparent blue envelope with the
atomic structure of bovine rhodopsin fitted in. There is good agreement between
the structures. The fit suggests that the lattice contacts may be made between
helix 8 (red) and helix 5 (orange). The linear arrangement in this lattice is
consistent with the in vivo alignment of squid rhodopsins for polarized
light discrimination.
Birkbeck
Crystallography EM and Image Processing Home Page
Birkbeck Crystallography
Home Page



