Saibil group, ISMB
Birkbeck College London
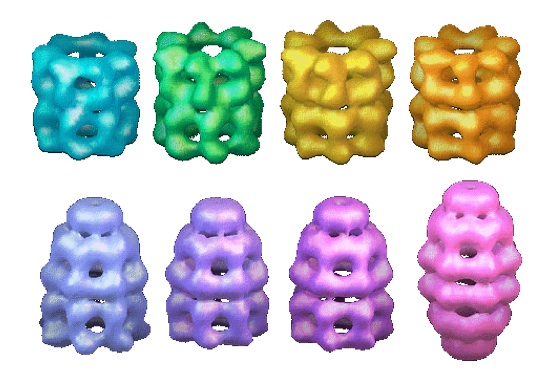

| Chaperones and protein
disaggregation Saibil group, ISMB Birkbeck College London |
 |
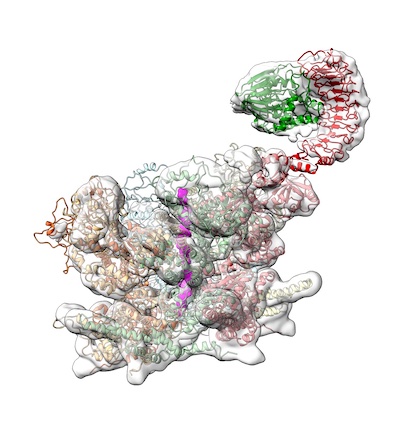 |
Structural
basis of ubiquitin-independent PP1 complex disassembly by
p97. van den Boom, J, Meyer, H & Saibil, HR (2022)
BioRxiv https://biorxiv.org/cgi/content/short/2022.06.24.497491v1 |
 |
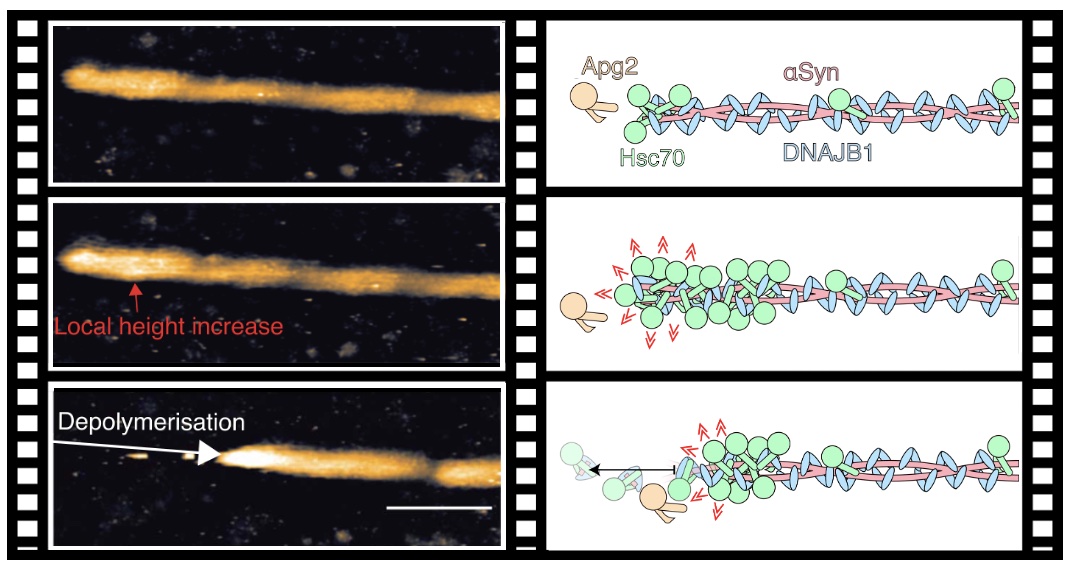 |
Cooperative
amyloid fibre binding and disassembly by the Hsp70
disaggregase. J.G. Beton, J Monistrol, A Wentink, EC
Johnston, AJ Roberts, B Bukau, BW Hoogenboom, HR Saibil
(2022) EMBO J https://doi.org/10.15252/embj.2021110410 |
|
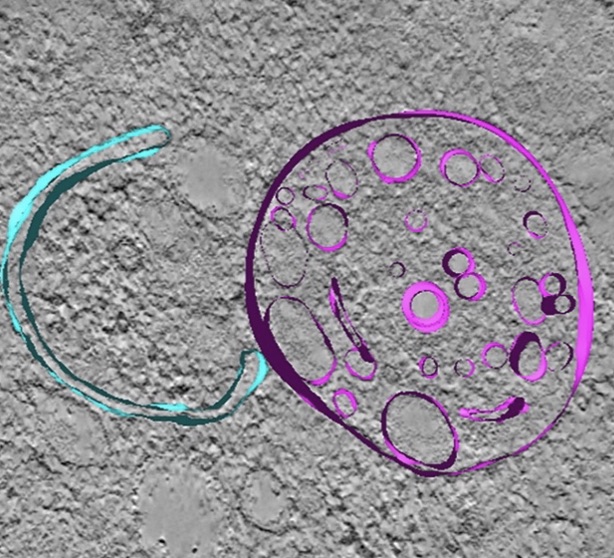 |
Correlative
light and electron microscopy suggests that mutant
huntingtin dysregulates the endolysosomal pathway in
presymptomatic Huntington's Disease. Zhou, Y, Peskett, TR,
Landles, C, Warner, JB, Sathasivam, K, Smith, EJ, Chen, S,
Wetzel R, Lashuel, HA, Bates, GP & Saibil, HR (2021) Acta
Neuropathologica Commun. 9:70 |
|
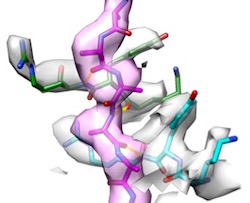 |
Two-step
activation mechanism of the ClpB disaggregase for
sequential substrate threading by the main ATPase motor.
Deville, C, Franke, K, Mogk, A, Bukau, B & Saibil, HR
(2019) Cell
Reports 27, 3433-3446. |
|
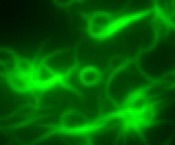 |
A liquid to solid phase
transition underlying pathological huntingtin exon1
aggregation. Peskett,
TR, Rau, F, O’Driscoll, J, Patani, R, Saibil, HR (2018)
Molec.
Cell, https://doi.org/10.1016/j.molcel.2018.04.007 |
|
|
Structural pathway
of regulated substrate transfer and threading through an
Hsp100 disaggregase. Deville, C, Carroni, M, Franke, KB,
Topf, M, Bukau, B, Mogk, A, & Saibil, HR (2017)
Science Advances 3:e1701726. |
||
 |
Human
Hsp70 disaggregase reverses Parkinson's-linked
alpha-synuclein amyloid fibrils. Gao, X, Carroni, M,
Nussbaum-Krammer, C, Mogk, A Nillegoda, NB, Szlachcic, A,
Guilbride, DL, Saibil, HR, Mayer MP & Bukau, B (2015)
Mol. Cell 59, 781-793. |
|
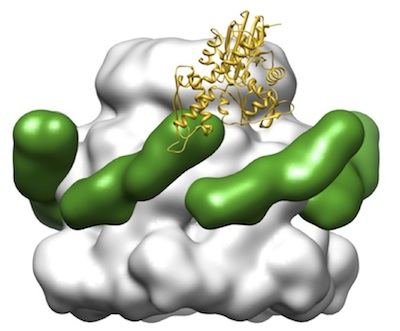 |
Head-to-tail interactions of
the coiled-coil domains regulate ClpB activity and
cooperation with Hsp70 in protein disaggregation.
Carroni, M, Kummer, E, Oguchi, Y, Wendler, P, Clare, DK,
Sinning, I, Kopp, J, Mogk, A, Bukau, B & Saibil, HR
(2014) eLife
3:e02481 |
|
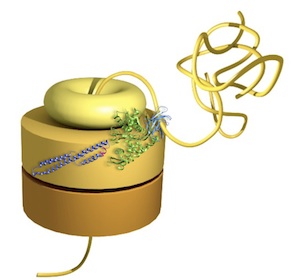 |
Chaperone machines for protein
folding, unfolding and disaggregation. Saibil, HR (2013)
Nature
Rev. Mol. Cell. Biol. 14, 630-642. |
|
 |
ATP-triggered conformational
changes delineate substrate-binding and -folding
mechanics of the GroEL chaperonin. Clare, DK, Vasishtan,
D, Stagg, S, Quispe, J, Farr, GW, Topf, M, Horwich, AL
& Saibil, HR (2012) Cell 149, 113-123. Movies |
|
 |
Newly
folded substrates inside the molecular cage of the HtrA
chaperone DegQ. Malet, H, Canellas, F, Sawa, J, Yan, J,
Thalassinos, K, Ehrmann, M, Clausen, T & Saibil, HR
(2012) Nature Struct. Mol. Biol. 19, 152-157.
|
|
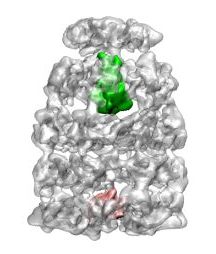 |
Chaperonin complex with a newly folded substrate protein encapsulated in the folding chamber. Clare, DK, Bakkes, P van Heerikhuizen, H, van der Vies, SM, & Saibil, HR (2009) Nature 457, 107-111. | |
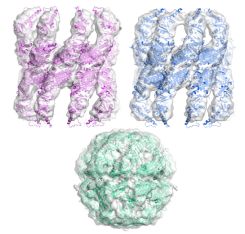 |
Multiple states of a
nucleotide-bound group 2 chaperonin. Clare, DK, Stagg,
S, Quispe, J, Farr, GW, Horwich, AL & Saibil, HR
(2008) Structure 16, 528-534, |
|
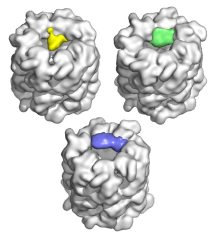 |
Topologies of a substrate protein bound to the chaperonin GroEL. Elad, N, Farr, GW, Clare, DK, Orlova, EV, Horwich, AL & Saibil, HR (2007) Mol. Cell 26, 415-426. | |
 |
Multiple distinct assemblies reveal conformational flexibility in the small heat shock protein Hsp26. White, HE, Orlova, EV, Chen, S, Wang, L, Ignatiou, A, Gowen, B, Stromer, T, Franzmann, TM, Haslbeck, M, Buchner, J & Saibil, HR (2006) Structure14, 1197-1204. | |
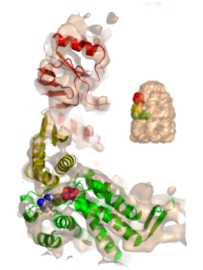 |
Allosteric signalling of ATP hydrolysis in GroEL-GroES complexes. Ranson, NA, Clare, DK, Farr, GW, Houldershaw, D, Horwich AL & Saibil, HR (2006) Nature Struct. Mol. Biol. 13, 147-152. | |
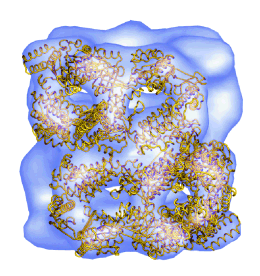 |
A mutant chaperonin with rearranged inter-ring electrostatic contacts and temperature-sensitive dissociation. Sewell, BT, Best, RB, Chen, S, Roseman, AM, Farr, GW, Horwich, AL & Saibil, HR (2004) Nature Struct. Mol. Biol. 11, 1128-1133. | |
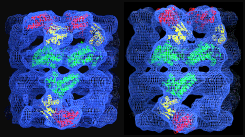 |
ATP-bound states of GroEL captured by cryo-electron microscopy, Ranson et al. (2001) Cell 107, 869-879. | |
 |
The chaperonin ATPase cycle:
mechanism of allosteric switching and movements of
substrate-binding domains in GroEL, Roseman et al.
(1996) Cell 87, 241-251. |
|
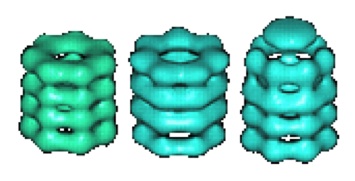 |
Location of a folding protein and shape changes in GroEL-GroES complexes imaged by cryo-electron microscopy, Chen et al. (1994) Nature 371, 261-264. |
Birkbeck EM group
Helen Saibil